I-AniI
Type
Wild-Type
Target Sequence
| -11 | -10 | -9 | -8 | -7 | -6 | -5 | -4 | -3 | -2 | -1 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 |
| C | T | G | A | G | G | A | G | G | T | T | T | C | T | C | T | G | T | A | A | A | G |
FASTA Sequence
MGDLTYAYLVGLFEGDGYFSITKKGKYLTYELGIELSIKDVQLIYKIKKILGIGIVSFRKRNEIEMVALRIRDKN
HLKSKILPIFEKYPMFSNKQYDYLRFRNALLSGIISLEDLPDYTRSDEPLNSIESIINTSYFSAWLVGFIEAEGC
FSVYKLNKDDDYLIASFDIAQRDGDILISAIRKYLSFTTKVYLDKTNCSKLKVTSVRSVENIIKFLQNAPVKLLG
NKKLQYKLWLKQLRKISRYSEKIKIPSNY
HLKSKILPIFEKYPMFSNKQYDYLRFRNALLSGIISLEDLPDYTRSDEPLNSIESIINTSYFSAWLVGFIEAEGC
FSVYKLNKDDDYLIASFDIAQRDGDILISAIRKYLSFTTKVYLDKTNCSKLKVTSVRSVENIIKFLQNAPVKLLG
NKKLQYKLWLKQLRKISRYSEKIKIPSNY
Crystal Structure
| PDB File |
| PyMOL File |
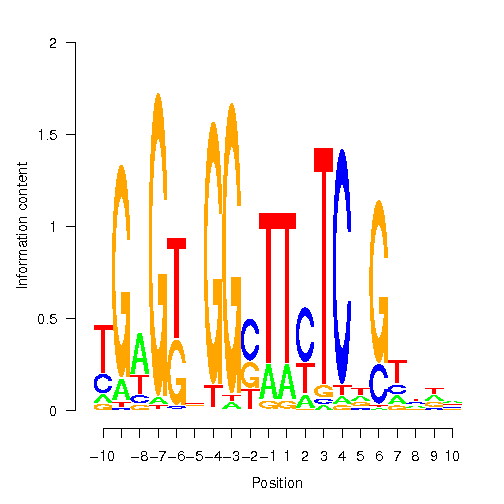
Position Weight Matrix
|
|
|
Amino Acid Contact Modules
| Module | Amino Acid Contact Modules (?) |
| -11 → -9: | Tyr 27 |
| -8 → -6: | Ser 20, Thr 22, Gly 25*, Thr 29, Glu 31, Ile 55, Ser 57, Arg 70, Arg 72 |
| -6 → -4: | Tyr 18, Ser 20, Gly 33, Glu 35, Ser 57, Arg 59*, Ala 68, Arg 70 |
| -5 → -3: | Tyr 18, Gly 33, Glu 35, Arg 59*, Glu 65*, Met 66, Ala 68, Arg 70 |
| +3 → +5: | Cys 150, Ser 152, Asp 168, Ala 170*, Asp 194*, Cys 198*, Lys 200 |
| +4 → +6: | Ser 152, Tyr 154, Ser 166, Asp 168, Lys 190*, Tyr 192, Asp 194*, Lys 202 |
| +6 → +8: | Tyr 154, Leu 156, Tyr 162, Ile 164*, Ser 166, Lys 190*, Thr 204 |
| +9 → +11: | Leu 156, Asp 159*, Asp 160*, Tyr 162 |
Key
| Experiment | |
| Crystal Structure | |
| Sequence Alignment | |
| * | Uncertain |
Specificity Changing Mutations
| Amino Acid Mutations (?) | Specificity Changes | Reference |
| K24N, T29K | -8A to -8G | 1 |
| L156Q, I164R, T204S | +8A to +8C | 1 |
| Y18W, E35K, R61Q | -3G to -3C | 1 |
| K24N, G25R, T29I | -9G to -9C | 1 |
| T29S, E31R, R70L | -6G to -6C | 1 |
| D168E | +5T to +5C | 1 |
| 1 | Thyme SB, Jarjour J, Takeuchi R, Havranek JJ, Ashworth J, Scharenberg AM, Stoddard BL, Baker D. (2009) Exploitation of binding energy for catalysis and design. Nature, 461, 1300-1304. |
Additional Information
Author: Barry StoddardNotes:
Date Added: 2012-01-29